Art and science have traditionally been treated as two separate disciplines. But scientists at Scripps Research show how the combination is allowing us to comprehend our microscopic world in unprecedented ways.
Structural and computational biologists at Scripps Research teamed up with talented molecular artists to produce the very first 3D model of an entire cell. The novel visualization of the bacterial cell Mycoplasma genitalium includes a detailed snapshot of its internal constituents and represents a new way to visualize cellular environments in health and medicine. The process was detailed in a recent paper in the Journal of Molecular Biology.
Akin to a list of ingredients, scientists in Scripps Research’s Center for Computational Structural Biology (CCSB) carefully curated the types and concentrations of the internal elements of the bacterial cell, from its DNA and RNA, to all of the functional proteins. Using simulation tools, scientists then assembled them to generate a unified picture of the biomolecular structure, building upon seminal work in 2012 by the Covert lab at Stanford University that had simulated the raw data for these intracellular components.
“The idea is to fill the gap between what we can see with normal light-microscopy and atomic-resolution techniques like X-ray crystallography or cryo-EM,” says Martina Maritan, PhD, a staff scientist in the lab of Professor David Goodsell, PhD. “Up until now there have not been techniques that can capture the range of scales from the molecular to the cellular level.”
Part of the reason why these complex models have not been possible is because scientists previously lacked adequately powered computer software. To overcome this problem, lead developer Ludovic Autin, PhD, borrowed technology developed by the gaming industry, using a game engine called Unity to simultaneously visualize millions of molecules and their precise interactions. They combined this technology with a suite of modelling programs called CellPACK, which builds upon the molecular color illustrations and augmented reality tools created by the CCSB, particularly the labs of Goodsell and Professor Arthur Olson, PhD.
“The technology is finally at a place where we can take advantage of the current flood of biological information and build models of this size and complexity,” says Goodsell.
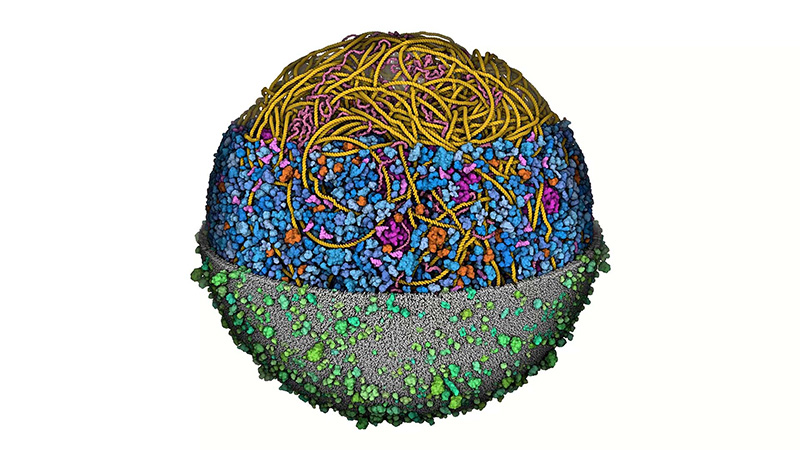
The model shows the outer cell membrane (grey/green), consisting of a layer of lipids and proteins that protect the interior of the cell from its external environment. Inside of the membrane depicts all the proteins in the cytoplasm (blue), the space where most cellular activities and chemical reactions occur. Nestled within this space are large complexes called ribosomes (magenta), which act as the cell’s protein-making machines. Beyond this layer is folded-up filaments of DNA (yellow) and mRNA (pink), the string of messenger code that contains the instruction manual for making each protein.
While the model is still unable to depict ions or water molecules, the prototype has already turned the classical notion of a cell on its head.
“Several high-school teachers and college professors who reached out to us remarked that they never realized how busy and crowded a cell was,” says Maritan. “The textbook idea of a cell as a simple sphere with a few objects floating in the middle is detached from reality.”
In addition to being a valuable educational tool for students, the model may also improve the way scientists create hypotheses and design experiments. Instead of thinking about certain molecules and their targets in isolation, models like these encourage researchers to think about the cellular components they study in the context of a complex environment.
Looking ahead, Goodsell’s team is interested in simulating other biological systems, including continued work on viruses such and HIV-1 and SARS-CoV-2, as well as more complex bacteria. They also are planning to model mitochondria, the energy factories of a cell that are implicated in various metabolic and neurodegenerative diseases. They hope these new visualization tools will be a major advancement in our basic understanding of cell biology, replacing abstract depictions with a fresh realist aesthetic.